April 30, 2026
Healthcare
Why Omics and Clinical Data Integration Remain a Challenge and How to Fix It
Key Takeaways
This article is about why that middle layer between both systems keeps breaking down and what a practical clinical data integration model looks like. Note that the root cause is almost always structural: separate systems, separate vocabularies, and separate governance. The article also explains how the silos form, reviews the multi-omics integration strategies worth knowing, and outlines the architectural pieces that make both datasets queryable together.
Neither is particularly powerful on its own for the hard questions in modern medicine. A variant of unknown significance in a gene stays a variant until a clinician can see how patients carrying it actually responded to a specific therapy. A transcriptomic subtype only becomes a clinical tool once it can be tied to survival curves.
A 2022 review in Computational and Structural Biotechnology Journal identified five translational objectives that depend directly on linking omics and clinical data:
But when connected effectively, a unified data repository delivers several benefits for precision medicine:
Multi-omics refers to the data from more than one molecular layer collected on the same sample. The most common layers are genomics (DNA sequence), transcriptomics (RNA expression), proteomics (proteins), epigenomics (DNA methylation, chromatin state), and metabolomics (small molecules).
Several of these layers now produce single-cell datasets, which add another layer of resolution. Kanda has covered the rise of that data type in detail in a separate article on single-cell atlas applications in life sciences.
Two core properties make multi-omics datasets hard to integrate even before clinical data enters the picture:
As Pythia’s primer on multi-omics integration points out, even agreeing on what counts as a “feature” across these layers is a non-trivial problem. Batch effects compound the issue: the same assay run on two different sequencers in two different weeks can produce measurable differences that have nothing to do with biology.
That volume interacts poorly with relational warehouses designed for transactional clinical records. This is why data-lake and data-mesh patterns have become relevant in healthcare; Kanda has compared those two architectures in detail in data lake vs. data mesh.
Multi-omics integration methods fall into roughly three families, and the choice matters before any architecture discussion because each strategy has different infrastructure implications downstream. A ScienceDirect review of integration strategies for machine learning and a more recent review in Briefings in Bioinformatics both group the options the same way.
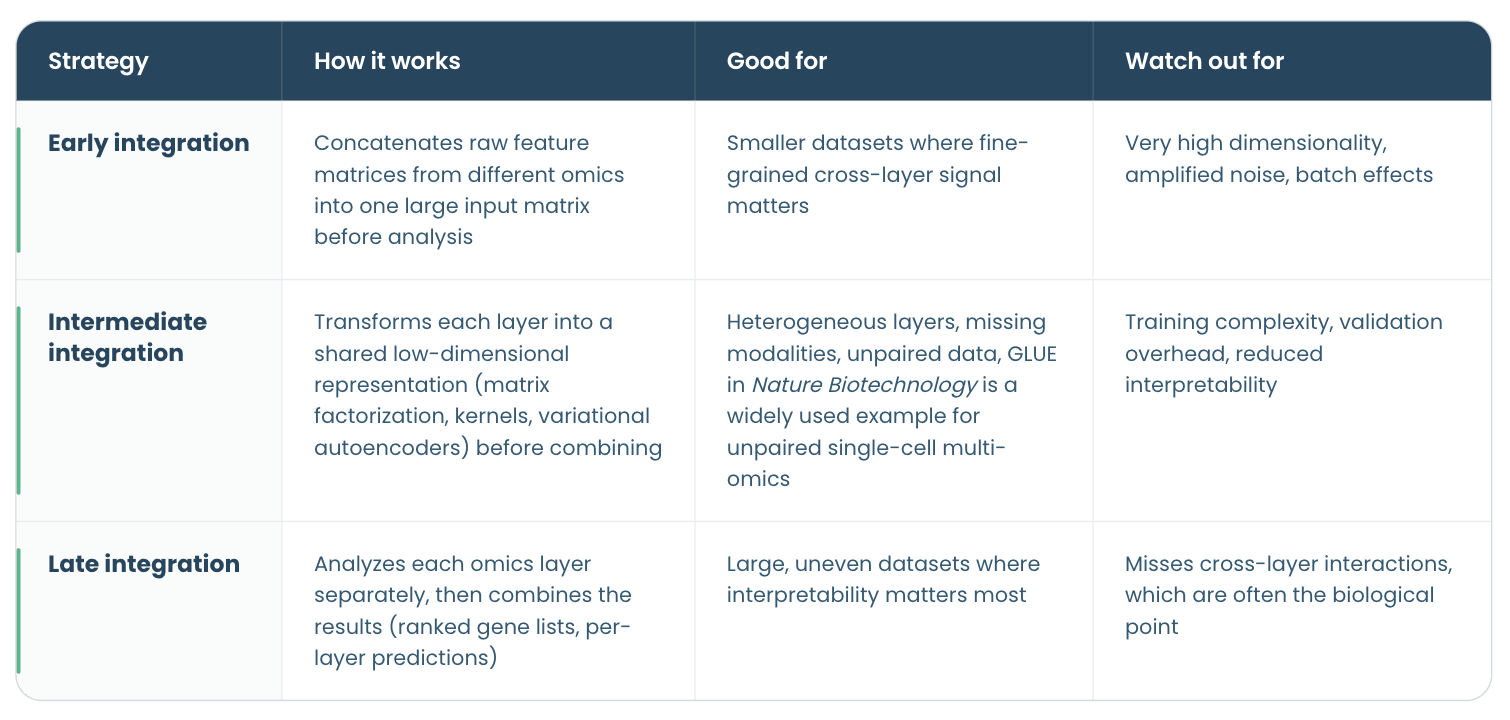 These strategies are orthogonal to how samples actually overlap across omics datasets. Teams often distinguish vertical integration (multiple modalities measured on the same patients), horizontal integration (the same modality compared across different cohorts), diagonal integration (different modalities on different patients with no paired samples), and mosaic integration (partial overlap, where some patients have some modalities and some do not). Which pattern your cohort fits into constrains the choice of early, intermediate, or late integration before any analysis begins.
These strategies are orthogonal to how samples actually overlap across omics datasets. Teams often distinguish vertical integration (multiple modalities measured on the same patients), horizontal integration (the same modality compared across different cohorts), diagonal integration (different modalities on different patients with no paired samples), and mosaic integration (partial overlap, where some patients have some modalities and some do not). Which pattern your cohort fits into constrains the choice of early, intermediate, or late integration before any analysis begins.
No single strategy dominates. The right one depends on the clinical question, the sample size, how complete the modalities are across patients, and how much interpretability the downstream user needs.
AI is also useful for imputing missing modalities, batch-effect correction, and unsupervised subtype discovery on cohorts too large to cluster by hand.
Where AI does not help—and where teams still overestimate its usefulness—is in the upstream plumbing: schema alignment, ontology mapping, and ID reconciliation. These are deterministic problems better solved by engineering than by models. Treat AI as a layer that sits on top of a clean integration pipeline, not as a substitute for building one.
On the semantic side, the single biggest lever is HL7 FHIR, together with SMART on FHIR for app-level integration. ONC’s 21st Century Cures Act Final Rule established FHIR Release 4 as the required API standard for certified health IT in the United States, and CMS built its Interoperability and Patient Access rule on the same foundation. Standardizing clinical terminology to SNOMED CT and LOINC, and tying omics annotations to shared data standards such as Gene Ontology, HPO, and HGVS, is what makes cross-system queries possible. Kanda’s EHR integration services practice builds on exactly these standards.
Underneath the semantic layer, the physical architecture has to keep clinical warehouses and bioinformatics pipelines from competing for resources. Clinical workloads require predictable, low-latency access to structured records, whereas omics workloads require bursty parallel compute on large immutable blobs. A cloud-native layout—object storage as the source of truth, compute clusters attached on demand, and open table formats such as Parquet, Iceberg, or Delta for tabular layers—lets each workload scale without dragging on the other. Decoupling storage from compute is what makes the standards above useful at scale rather than just in pilots.
The clinical research that makes precision medicine work depends on this layer being reliable, repeatable, and shared. Organizations that invest in it early stop losing months per project to data wrangling and redirect that time to the science itself.
- Omics and clinical data often remain separate, making them harder to use together than they should be.
- The main challenge is usually not the data itself, but the mismatched formats, siloed systems, and disconnected teams surrounding it
- Addressing that requires robust infrastructure, shared standards, and workflows that hold up without constant manual intervention.
- AI can manage analytical complexity, but it is no substitute for sound engineering and clear governance.
- When integration is done well, researchers spend less time wrestling with their data and more time drawing meaning from it.
This article is about why that middle layer between both systems keeps breaking down and what a practical clinical data integration model looks like. Note that the root cause is almost always structural: separate systems, separate vocabularies, and separate governance. The article also explains how the silos form, reviews the multi-omics integration strategies worth knowing, and outlines the architectural pieces that make both datasets queryable together.
How the Fusion of Omics and Clinical Data Changes Precision Medicine
Multi-omics measurements describe the molecular state of a patient across multiple layers of biological systems: what their genome looks like, what their cells are transcribing, what proteins are present, and which metabolites are circulating. Clinical data describes the medical state and treatment history, including symptoms, diagnoses, and outcomes over time.Neither is particularly powerful on its own for the hard questions in modern medicine. A variant of unknown significance in a gene stays a variant until a clinician can see how patients carrying it actually responded to a specific therapy. A transcriptomic subtype only becomes a clinical tool once it can be tied to survival curves.
A 2022 review in Computational and Structural Biotechnology Journal identified five translational objectives that depend directly on linking omics and clinical data:
- Detecting disease-associated molecular patterns
- Disease subtyping
- Diagnosis and prognosis
- Drug-response prediction
- Understanding biological mechanisms.
But when connected effectively, a unified data repository delivers several benefits for precision medicine:
- A more holistic view of diseases, enabling better-targeted therapy.
- Improved therapeutic success through effective treatment optimization.
- Better disease classification, supported by both omics and clinical data.
Understanding Multi-Omics Data
To fully understand the challenge of uniting omics and clinical data, it is helpful to know what multi-omics data is and what makes it distinctive in terms of management, storage, and governance.Multi-omics refers to the data from more than one molecular layer collected on the same sample. The most common layers are genomics (DNA sequence), transcriptomics (RNA expression), proteomics (proteins), epigenomics (DNA methylation, chromatin state), and metabolomics (small molecules).
Several of these layers now produce single-cell datasets, which add another layer of resolution. Kanda has covered the rise of that data type in detail in a separate article on single-cell atlas applications in life sciences.
Two core properties make multi-omics datasets hard to integrate even before clinical data enters the picture:
Data heterogeneity
Each omics layer produces different data types and has its own measurement technology, file format, noise profile, and statistical structure. Genomic variants arrive as VCF files. Transcriptomics produces gene-expression matrices in HDF5 or MTX. Proteomics data outputs peak lists or spectra. Metabolomics data is often unaligned across vendors.As Pythia’s primer on multi-omics integration points out, even agreeing on what counts as a “feature” across these layers is a non-trivial problem. Batch effects compound the issue: the same assay run on two different sequencers in two different weeks can produce measurable differences that have nothing to do with biology.
Data volume
A whole-genome sequence produces roughly 100–200 GB of raw reads per patient. A typical single-cell transcriptomics dataset for a few thousand cells runs into tens of gigabytes. Imaging-based assays add more. An institution running routine multi-omics studies across a few thousand patients a year faces petabyte-scale storage before clinical data is even attached.That volume interacts poorly with relational warehouses designed for transactional clinical records. This is why data-lake and data-mesh patterns have become relevant in healthcare; Kanda has compared those two architectures in detail in data lake vs. data mesh.
Why Healthcare Data Silos Form Between Clinical and Omics Environments
Healthcare data silos usually form because clinical IT and research IT are built for different primary users—clinicians for one, scientists for the other—and historically answered to different regulatory regimes.. A few patterns explain most of the breakage.Missing and corrupted data
Clinical records are full of gaps, often the result of human error. Missed measurements, lab values entered as free text, and medications reconciled inconsistently at discharge are common examples. Omics pipelines fail when upstream sequencing runs miss coverage thresholds or when sample contamination slips past initial QC. Integrating clinical and omics data exposes gaps on both sides at once, making the combined dataset harder to trust and more difficult to manage.Lack of computational power
The infrastructure hosting EHRs is tuned for transactional workloads and low-latency lookups, whereas omics analyses need distributed compute, elastic storage, and GPU access. Running a variant-calling pipeline on a hospital data warehouse is often architecturally impossible without a parallel research environment.Miscommunication between outdated systems
Many hospitals still run clinical systems that predate modern APIs. A bioinformatics pipeline expecting structured, machine-readable patient data frequently receives CSV extracts from an HL7 v2 interface engine, or PDFs from a pathology LIMS. Each boundary between systems introduces a translation step that can drift out of sync with the source system.Human error from manual entry
Building on the point above, wherever integration requires manual data reconciliation and mapping, sample IDs to patient IDs, tissue codes to ICD-10 diagnoses, consent records to research cohorts, mistakes compound silently. A PhD student reconciling IDs in Excel is the modal failure mode of modern translational research.Lack of semantic interoperability
This is the issue most teams underestimate. The same cancer diagnosis can appear in very different forms across systems: as a billing code in the EHR, as a standardized clinical term, or simply as free text in physician notes. Meanwhile, a bioinformatics pipeline classifying tumor subtypes, meanwhile, uses its own vocabulary, such as Gene Ontology terms, pathway IDs, or HGVS variant nomenclature. Joining those representations requires ontology-mapping layers that most institutions never formally build.Consent and regulatory boundaries
Research data collected under an IRB protocol and clinical data generated as part of routine care are governed by different rules, and are often stored on different systems by design. Mixing them requires explicit data-use agreements, re-consent workflows, or honest-broker arrangements. Architectures that ignore these boundaries tend to either leak data that shouldn’t move or block legitimate research that should.Three Multi-Omics Integration Strategies
With a clearer picture of what multi-omics data is and why pipelines break, the next question is how to fix it and integrate everything into a frictionless workflow. Several strategies are available.Multi-omics integration methods fall into roughly three families, and the choice matters before any architecture discussion because each strategy has different infrastructure implications downstream. A ScienceDirect review of integration strategies for machine learning and a more recent review in Briefings in Bioinformatics both group the options the same way.
 These strategies are orthogonal to how samples actually overlap across omics datasets. Teams often distinguish vertical integration (multiple modalities measured on the same patients), horizontal integration (the same modality compared across different cohorts), diagonal integration (different modalities on different patients with no paired samples), and mosaic integration (partial overlap, where some patients have some modalities and some do not). Which pattern your cohort fits into constrains the choice of early, intermediate, or late integration before any analysis begins.
These strategies are orthogonal to how samples actually overlap across omics datasets. Teams often distinguish vertical integration (multiple modalities measured on the same patients), horizontal integration (the same modality compared across different cohorts), diagonal integration (different modalities on different patients with no paired samples), and mosaic integration (partial overlap, where some patients have some modalities and some do not). Which pattern your cohort fits into constrains the choice of early, intermediate, or late integration before any analysis begins.No single strategy dominates. The right one depends on the clinical question, the sample size, how complete the modalities are across patients, and how much interpretability the downstream user needs.
How to Achieve Reliable Clinical Data Integration
Picking an integration strategy is only half the job. The other half is building the architecture that makes it work reliably at scale. The patterns below work across most enterprise healthcare systems and life sciences environments.1. Deploy AI where it earns its place
AI and machine learning pay off in a few specific parts of the integration stack, not across the board. The clearest wins are in the intermediate-integration family described above. Methods like variational autoencoders and graph-based models often perform better when data layers are heterogeneous and incomplete.AI is also useful for imputing missing modalities, batch-effect correction, and unsupervised subtype discovery on cohorts too large to cluster by hand.
Where AI does not help—and where teams still overestimate its usefulness—is in the upstream plumbing: schema alignment, ontology mapping, and ID reconciliation. These are deterministic problems better solved by engineering than by models. Treat AI as a layer that sits on top of a clean integration pipeline, not as a substitute for building one.
2. Standardize biomedical data integration
Standardization in a biomedical data integration platform works on two levels that reinforce each other: the semantic layer (how the data is described) and the physical layer (how it is stored and served).On the semantic side, the single biggest lever is HL7 FHIR, together with SMART on FHIR for app-level integration. ONC’s 21st Century Cures Act Final Rule established FHIR Release 4 as the required API standard for certified health IT in the United States, and CMS built its Interoperability and Patient Access rule on the same foundation. Standardizing clinical terminology to SNOMED CT and LOINC, and tying omics annotations to shared data standards such as Gene Ontology, HPO, and HGVS, is what makes cross-system queries possible. Kanda’s EHR integration services practice builds on exactly these standards.
Underneath the semantic layer, the physical architecture has to keep clinical warehouses and bioinformatics pipelines from competing for resources. Clinical workloads require predictable, low-latency access to structured records, whereas omics workloads require bursty parallel compute on large immutable blobs. A cloud-native layout—object storage as the source of truth, compute clusters attached on demand, and open table formats such as Parquet, Iceberg, or Delta for tabular layers—lets each workload scale without dragging on the other. Decoupling storage from compute is what makes the standards above useful at scale rather than just in pilots.
3. Build automated pipelines for omics
Treat each stage of the pipeline – ingestion, transformation, harmonization, and serving – as an independent service with its own inputs, outputs, and tests. A good omics data integration pipeline ingests raw files into a landing zone, runs validated bioinformatics workflows (GATK, nf-core, Bioconductor tools) in isolated compute environments, harmonizes outputs against a shared patient and sample vocabulary, and serves the integrated data to both research and clinical consumers through well-documented APIs. Orchestrators such as Airflow, Dagster, or Nextflow make dependencies explicit and recoverable. Done well, this is what turns a one-off analysis into a reusable clinical data integration model rather than another bespoke extract.4. Audit your existing data management infrastructure first
Before rebuilding anything, map what you already have. Starting from your core data sources, follow a handful of patient cohorts end to end, from the sequencing lab to the clinical dashboard, and document where each step lives, who owns it, how long it takes, and where the handoffs are manual. Benchmark latency and failure rates. Most teams find two or three choke points that account for most of the pain, and addressing those is often more cost-effective than a full re-platforming. Kanda’s best practices guide for medtech system integration walks through a similar audit approach for device data.5. Bake in security and observability from day one
HIPAA, GDPR, and institution-specific IRB requirements are not a regulatory compliance afterthought in a biomedical data integration platform, they are design constraints. Role-based access control, column-level encryption for PHI and genotype data, immutable audit logs, and consent enforcement should be part of the initial architecture. Observability is the mirror image: lineage tracking from raw files to derived features, data-quality checks at every transformation stage, and alerts that catch silent pipeline drift before a downstream analysis ships wrong results.When to Bring in a HealthTech Development Partner
- Internal teams can go a long way on this kind of work, but a few situations make bringing in an external partner pay off quickly:
- The organization has strong clinical-informatics talent but limited bioinformatics data-engineering experience (or the reverse), and the integration work sits squarely between them.
- Regulatory timelines, 21st Century Cures Act enforcement, EU HTA, MDR, IVDR, are compressing the runway, and the team needs additional hands on deck now.
- A working prototype needs to scale to multi-site or multi-cohort use, and the team that built it isn’t set up to operate production systems.
- The team wants an outside view from someone who has seen the same pattern across multiple healthcare and life sciences organizations, and can point out which shortcuts tend to backfire.
How Kanda Can Help
Kanda has been building software for healthcare, life sciences, and pharmaceutical companies for more than 30 years. For clinical and multi-omics data integration specifically, the work usually falls into a few categories:- Designing a practical data management and analytics strategy that covers both clinical and research data, with clear ownership and governance.
- Implementing EHR integration against FHIR, HL7 v2, and custom interfaces, including semantic mapping to SNOMED CT, LOINC, and ICD-10.
- Building cloud-native data platforms that decouple storage from compute and use data lake or data mesh patterns appropriate to the organization’s scale and team shape.
- Standing up automated bioinformatics pipelines with lineage, reproducibility, and clean handoffs to downstream clinical and research consumers.
- Adding HIPAA-grade security, consent enforcement, and observability to existing integration architectures without starting over.
Conclusion
The gap between omics and clinical data is real and costly, but it is not mysterious. Systems were built for different users, governed under different rules, and run on infrastructure optimized for different workloads.The clinical research that makes precision medicine work depends on this layer being reliable, repeatable, and shared. Organizations that invest in it early stop losing months per project to data wrangling and redirect that time to the science itself.
Related Articles

CMS Prior Authorization Rule 2026: What Health Systems and Payers Actually Need to Build by October 2027
Key Takeaways The 2026 CMS Prior Authorization (CMS-0062-P) rule extends electronic prior authorization requirements to drugs for the first time, closing a major gap left by the 2024 rule. The hard compliance deadline is October 1, 2027 - by that date, systems must be live, not in development or testing. Compliance requires three major technical…Learn More
Unlocking Actionable Insights from Medical Data with Clinical NLP
Key Takeaways Clinical NLP extracts useful details from notes, reports, and other free-text records so teams can work with them. The clearest use cases include coding support, trial matching, safety monitoring, registry work, and population health. The output is only useful if it connects to the standards healthcare teams already use: SNOMED CT, RxNorm, LOINC,…Learn More
AI Copilots in Healthcare: Minimizing Diagnostic Risks With Smart Virtual Assistants
Key Takeaways AI copilots act as always-on virtual assistants managing documentation, triage, and decision support so clinicians can focus on patients. With 12 million diagnostic errors and up to 80,000 deaths annually, AI copilots are built to catch what overloaded clinicians miss, not replace them. Dragon Copilot saves clinicians around 5 minutes per encounter with…Learn More
Change Management in Healthcare Software: The Missing Piece of Digital Transformation
Key Takeaways Nearly two-thirds of healthcare change projects fail, and the root cause is almost always people and process, not technology. Clinician resistance is a rational response to workflow disruption, not a personality problem. Plan for it. Change management is a parallel workstream that runs alongside software development from day one, not a training session…Learn More

